Inferring intrinsic neural timescales using optimal control theory
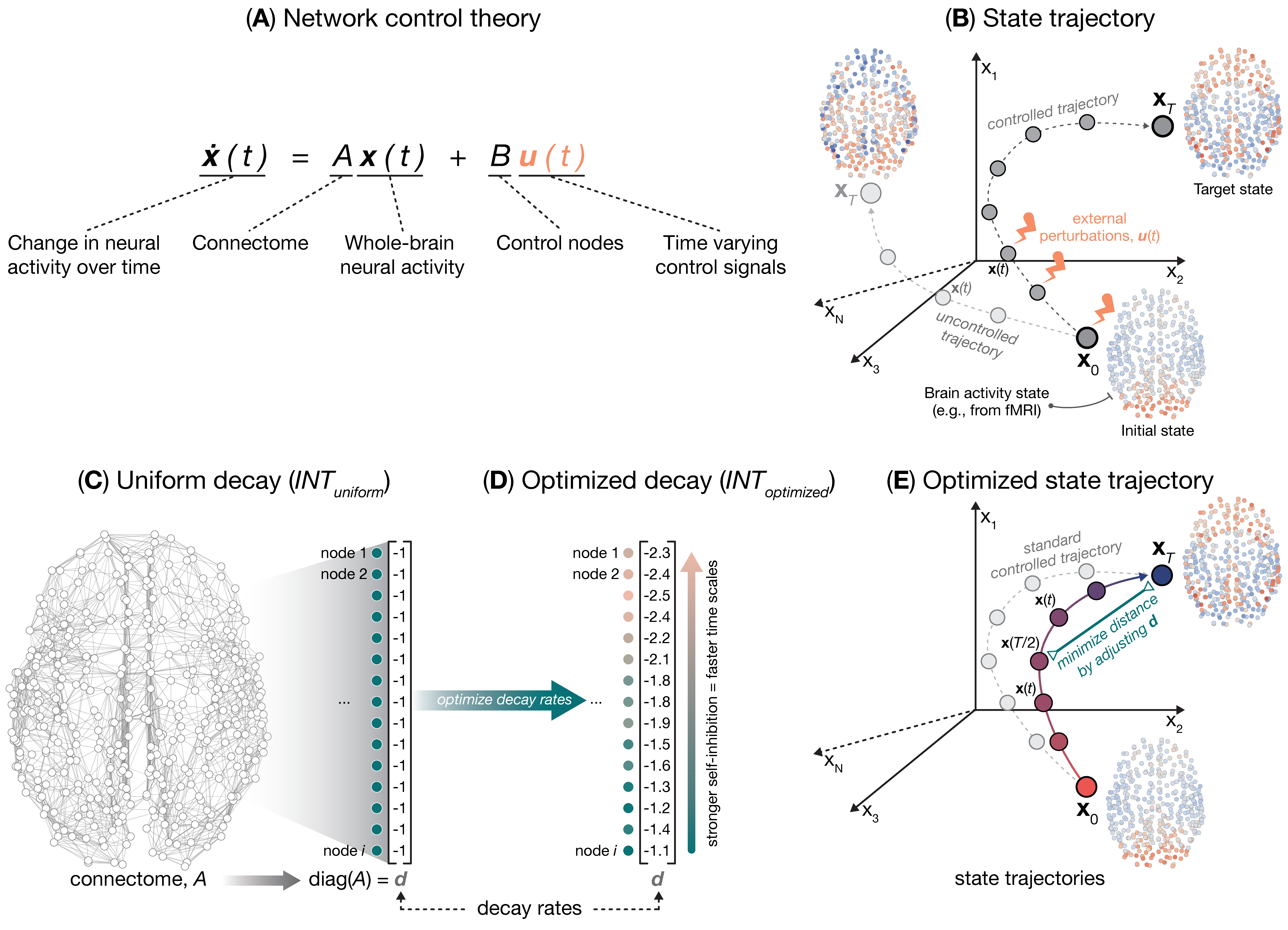
The temporal evolution of whole-brain activity is contingent upon complex interactions within and between brain regions that are mediated by neurobiology and connectivity, respectively. Here, we provide a framework for studying these relationships that uses network control theory (NCT) to estimate regions’ intrinsic neural timescales (INTs). Our approach broadens the range of dynamics supported by the connectome and improves the alignment between the brain’s connectivity and its traversal through state-space. We find that our model-based INTs correlate with INTs measured empirically from functional neuroimaging data, neurobiological measures of gene expression and cell-type densities, as well as measures of cognition. We demonstrate consistent results across multiple datasets and species. Finally, we show that our model-based INTs enable the efficient control of brain states from fewer brain regions. Our results provide a flexible quantitative framework that more accurately captures the interplay between brain structure, function, and intrinsic dynamics with greater biophysical realism.
Project Leads
Jason Z. Kim
Faculty Leads
Linden Parkes
Collaborators
Richard F. Betzel, Ahmad Beyh, Amber Howell, Amy Kuceyeski, Bart Larsen, Caio Seguin, Xi-Han Zhang, Avram Holmes
Project Start Date
August 2023
Current Project Status
Published in Nature Communications (2025) as Inferring intrinsic neural timescales using optimal control theory
Datasets
Human Connectome Project Young Adult (HCP-YA), Microstructure-Informed Connectomics (MICA-MICs), Allen Mouse Brain Connectivity Atlas, Allen Human Brain Atlas
Github Repository
https://github.com/LindenParkesLab/nct_xr
Path to Data on Filesystem
n/a
Publication DOI
https://doi.org/10.1038/s41467-025-66542-w
Conference Presentations
- Oral presentation at the The Organization for Human Brain Mapping Annual Meeting, June 2025. Inferring intrinsic neural timescales using optimal control theory.